Research
SNARE facilitates exploration of data described in Brase, et al. (2023) Single-nucleus RNA-sequencing of autosomal dominant Alzheimer disease and risk variant carriers .
Authors
Logan Brase, Shih-Feng You, Jorge del Aguila, Yaoyi Dai, Brenna C Novotny, Carolina Soriano-Tarraga, Taitea Dykstra, Maria Victoria Fernandez, John P Budde, Kristy Bergmann, John C Morris, Randall J Bateman, Richard J Perrin, Eric McDade, Chengjie Xiong, Alison Goate, Martin Farlow, Jasmeer P. Chhatwal, Peter Schofield, Helena Chui, Dominantly Inherited Alzheimer Network (DIAN), Greg Sutherland, Jonathan Kipnis, Celeste M Karch, Bruno A Benitez, Carlos Cruchaga, Oscar Harari
Background
Alzheimer Disease (AD) has substantial genetic, molecular, and cellular heterogeneity associated with its etiology. Much of our current understanding of the main AD molecular events associated with the amyloid hypothesis (APP, PSEN1 and PSEN2) and neuroinflammation (TREM2) is based on genetic studies. However, the functional genes, downstream transcriptional ramifications and the cell-type-specific effects of these genetic factors remains poorly understood. Understanding these effects can point us to cellular processes involved in AD progression and uncover potential therapeutic targets.
Methods
We applied a genetic-based approach to our sample selection; our cohort includes carriers of AD pathogenic mutations (APP, PSEN1), risk variants in TREM2, and the resilient variant (rs1582763) in the MS4A cluster associated with cerebrospinal fluid (CSF) soluble TREM2 levels. We performed single nucleus RNA-sequencing (snRNA-seq) of 1,102,459 nuclei from the human parietal cortex of these carriers. Following initial unbiased clustering and cell-type annotation, we performed deep subclustering analysis per cell type to identify unique cellular transcriptional states associated with these genetic variants. We identified differentially expressed genes between cell states and genetic variant carriers/controls, and performed differential cell proportion analyses to determine key differences among these carriers. We analyzed single nucleus data from human dorsolateral prefrontal cortex and mice models to replicate the enrichment of unique cell states in genetic variant carriers. Finally, we leveraged these cell-state differential expression results to link genes in AD GWAS loci to their functional cell types.
Findings
We identified cell-specific expression states influenced by AD genetic factors for neurons and glia. Autosomal dominant AD (ADAD) brains exhibited unique transcriptional states in all cell types. TREM2 variant carrier brains were also enriched in specific microglia and oligodendrocyte subpopulations. We mapped AD GWAS genes to their potential functional cell types, and some, including PLCG2 and SORL1, are expressed in a broader range of brain cell types than previously reported.
Interpretation
AD pathogenic, risk or resilient variants are sufficient to alter the transcriptional and cellular landscape of human brains. Overall, our results suggest that the genetic architecture contributes to the cortical cellular heterogeneity associated with disease status, which is a critical factor to consider when designing drug trials and selecting the treatment program for AD patients. Our findings suggest that integrating genetic and single-cell molecular data facilitates our understanding of the heterogeneity of pathways, biological processes and cell types modulated by genetic risk factors for AD.
Usage
When visualizing gene expression, we recommend setting the clip percentile range to 0-99% to avoid desaturation:

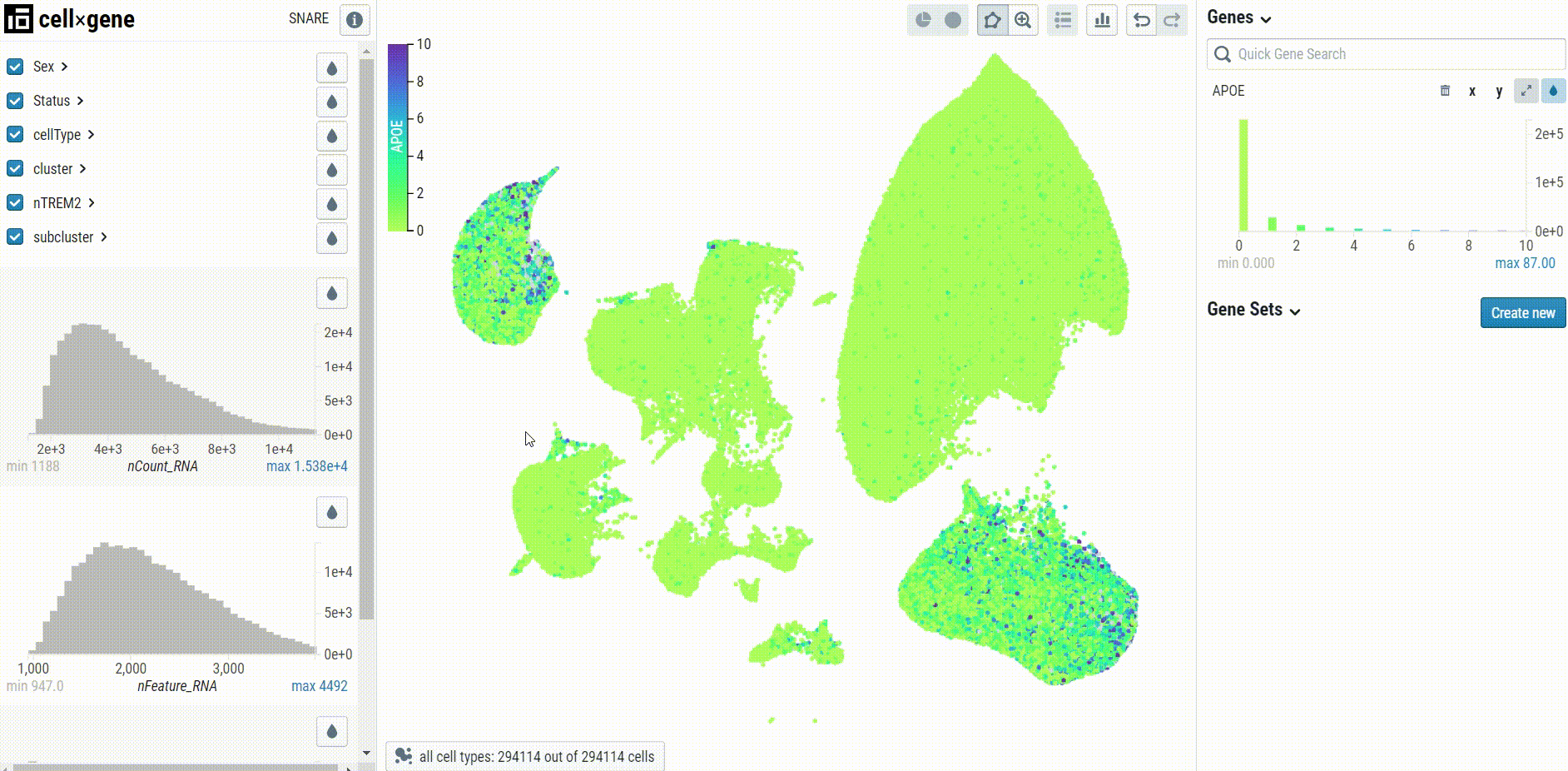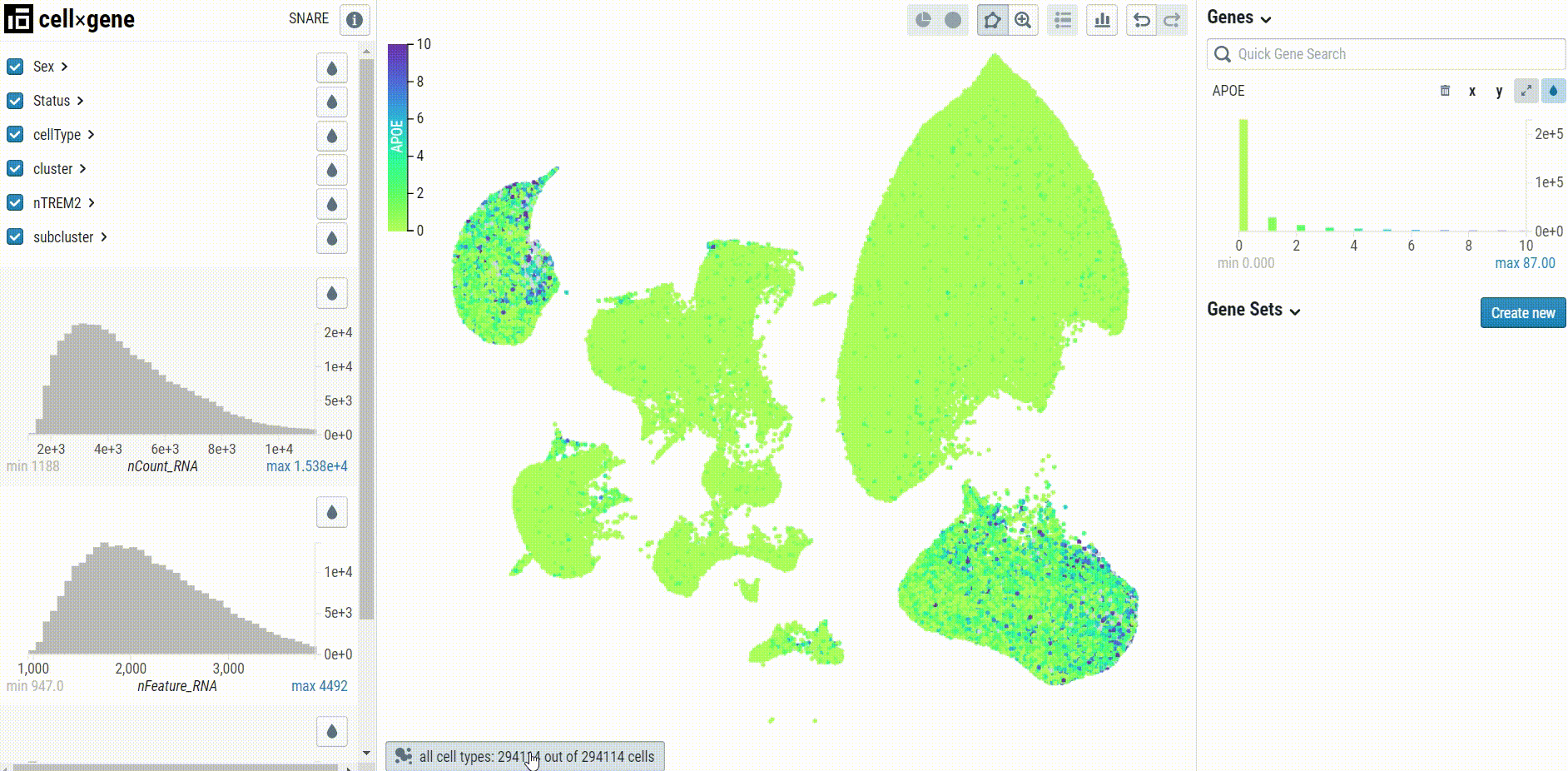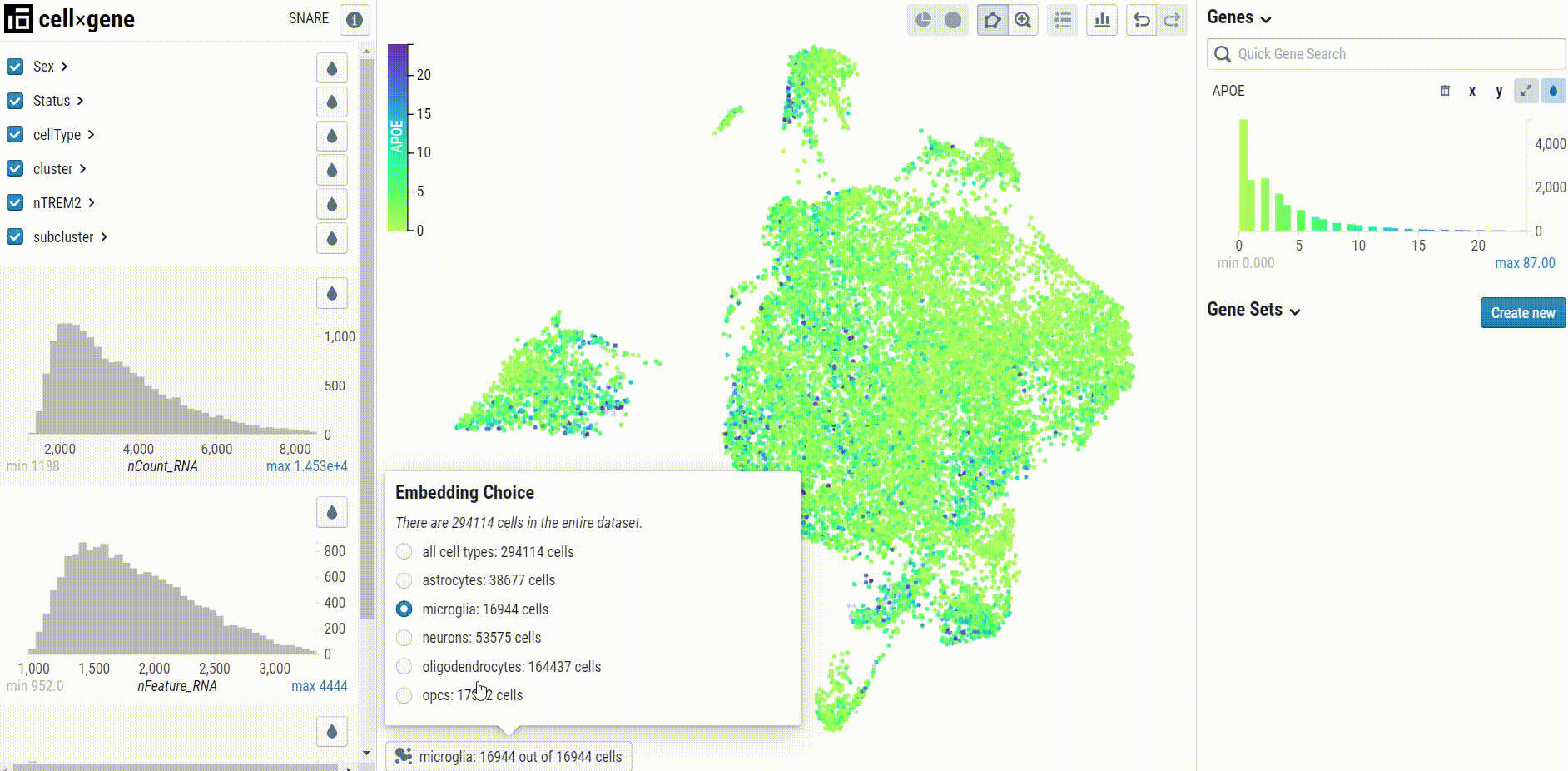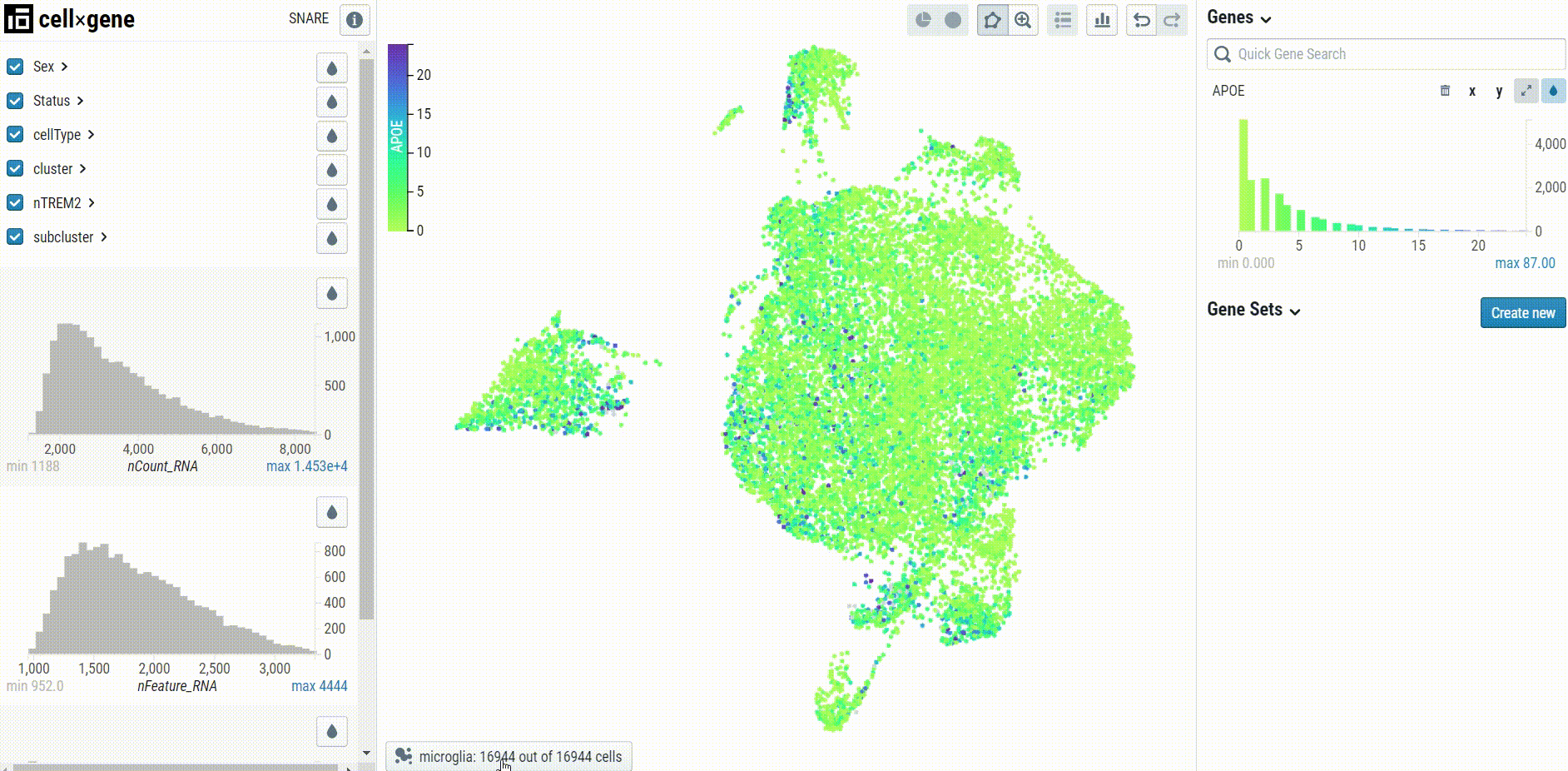
Cell type-specific UMAP plots may be explored by selecting the embedding in the bottom left corner of the UMAP pane:
The SNARE browser was built with cellxgene from the Chan Zuckerberg Initiative. See their documentation for more details on exploring data.
Go to SNARE